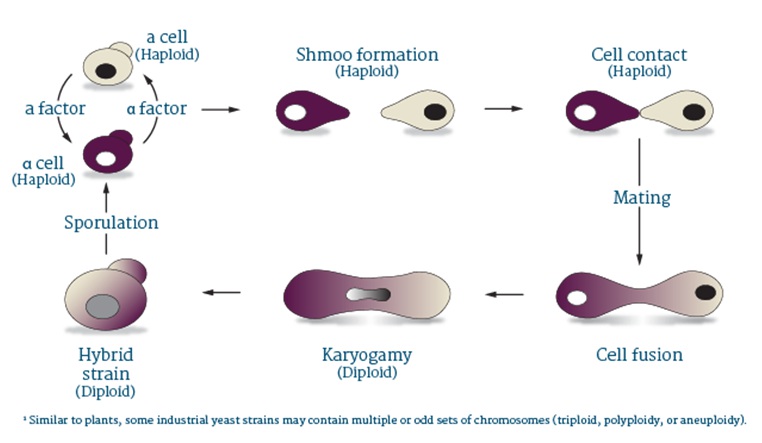
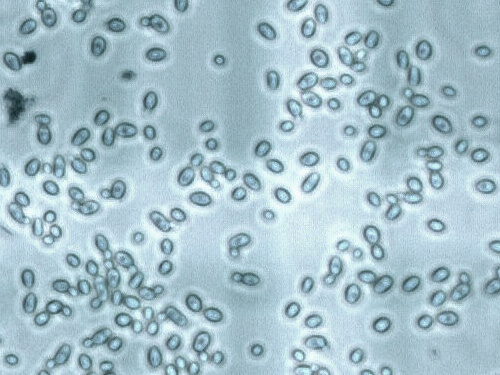
About Saccharomyces eubayanus
A Little History of Lager Yeast
For millennia humans have cultivated the yeast Saccharomyces cerevisiae for its use in making bread, beer, wine, cider and mead. In contrast to today’s pitchable packages of pure, clonal, quality-controlled yeast strains, the yeast cultivated by brewers prior to the late 19th century also contained other microorganisms. As the nutrients in wort are quickly consumed by S. cerevisiae to produce ethanol and lower the pH, an environment that favors S. cerevisiae growth over its rival species ensures it remains the dominant species in the yeast culture. Yet things began to change in the Middle Ages when brewing during the summer months in Bavaria was prohibited, forcing brewers to ferment and store beer at colder temperatures (1). S. cerevisiae growth is slow at colder temperatures, thereby presenting an opportunity for other microorganisms to dominate the culture. This opportunity was seized by the mysterious arrival of the yeast S. eubayanus, and thus began a brewing revolution.
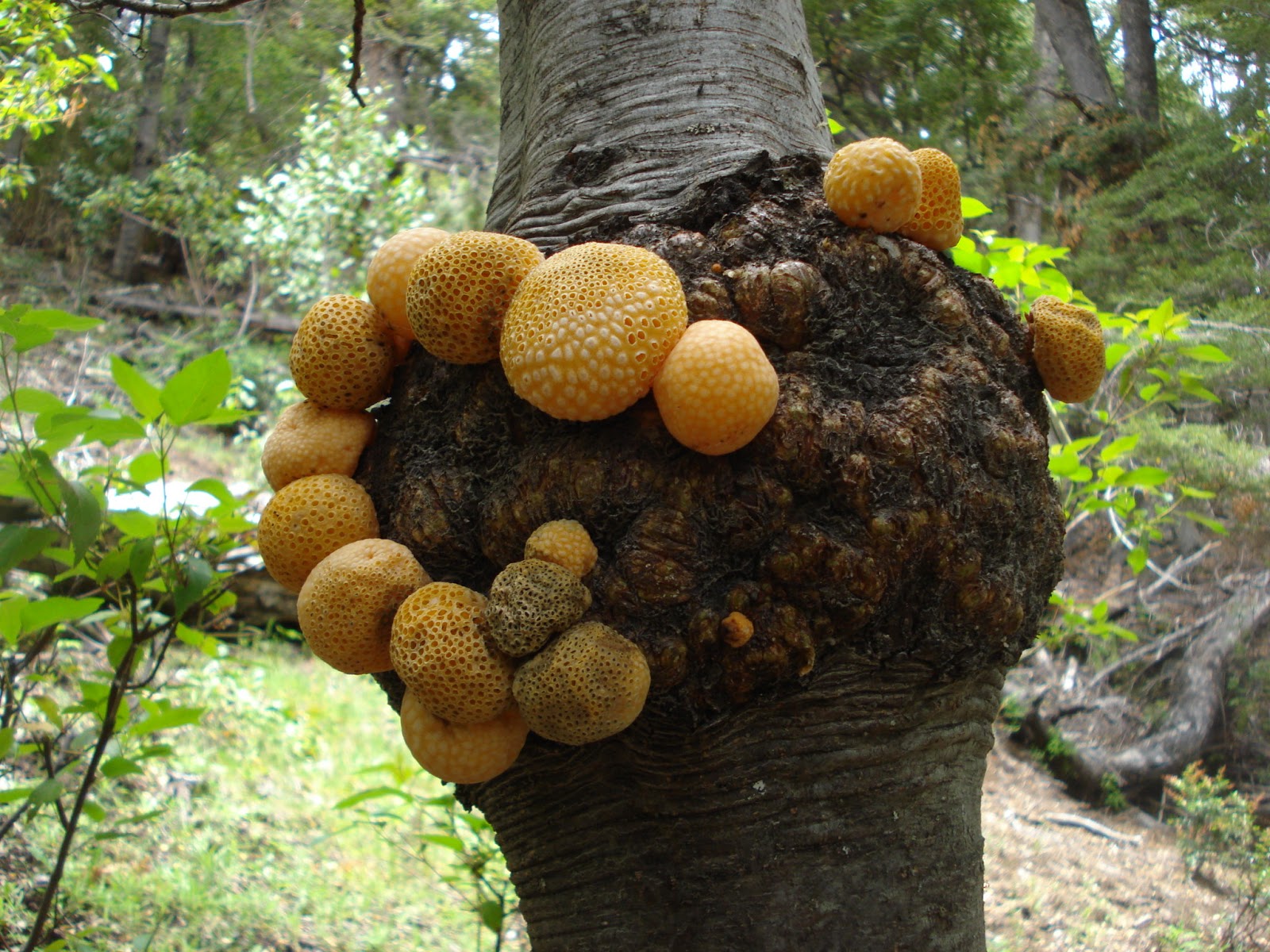
S. eubayanus is a cold-tolerant yeast discovered in 2011 on trees in Patagonia (2) and was subsequently found in China and Wisconsin (3, 4). It’s uncertain how it got into Bavarian beer, but through the process of selecting cold-tolerant yeast, brewers inadvertently created lager yeast S. pastorianus resulting from the hybridization of S. cerevisiae and S. eubayanus. The daughter yeast had two genomes, allowing it to compete with both S. cerevisiae and S. eubayanus. As the yeast began dividing and adapting to its environment, unnecessary or redundant genes were removed by chromosomal deletion and/or rearrangement. S. pastorianus refined its genome to become the superior yeast and eventually out-competed both S. cerevisiae and S. eubayanus to become the dominant yeast in lager cultures.
Yeast Hybrids
Hybrid species of Saccharomyces are not uncommon in the production of fermented beverages, as many hybrids have been identified in the wine- and cider-making industries (5). Although it is quite likely that multiple, independent hybridization events gave rise to genetically diverse strains of S. pastorianus, there is remarkably very little genetic diversity. This is most likely due to the use of pure strains of yeast set forth by Pasteur and Hansen in the late 19th century. Hansen isolated the first lager strain at the Carlsberg Laboratories (S. carlsbergenis), and this pure strain was freely distributed to other breweries. At the turn of the 20th century, Paul Lindner (also at the Carlsberg Laboratories) isolated two individual S. pastorianus strains (6, 7). These two strains (Saaz and Frohberg strains, named after the location of their respective breweries) subsequently replaced mixed lager cultures, and all modern day lager strains are derived from these early isolates. The difference in fermentation between Frohberg and Saaz stains has long been recognized. Saaz stains typically ferment quicker at colder temperatures than the Frohberg strains, while the latter tends to attenuate much more.
Genetic sequencing of Frohberg and Saaz strains shows each group arose from two independent hybridization events (8-11). After S. cerevisiae and S. eubayanus hybridized, the resulting yeast cell contained multiple copies of each chromosome. Over time, resulting from selective pressure from brewers, duplicate chromosomes either chimerized via homologous recombination or they were simply deleted.
S. eubayanus Fermentation Profile
The chromosomal makeup of Saaz strains are triploid (one copy of the S. cerevisiae genome, two copies of the S. eubayanus genome), and Frohberg stains are tetraploid (two copies of both S. cerevisiae and S. eubayanus genomes) (8-11). A recent study comparing fermentations by Frohberg and Saaz strains and S. eubayanus showed that Saaz strains performed very similar to S. eubayanus, consistent with there being more S. eubayanus chromosomes in the genome. Whereas the Frohberg strains were highly attenuative, both S. eubayanus and the Saaz strains had low apparent attenuations (approximately 65%). The Saaz stains and S. eubayanus did not ferment very well at warm temperatures, and are much more cold-tolerant than the Frohberg strains (12).
The lower attenuation seen in S. eubayanus is due to a deficiency in transporting maltotriose into the cell. This was previously shown in the Saaz strains as well (12). This study suggested that the inability to ferment maltotriose was not due to an inability of the cell to convert maltotriose to ethanol, but due to a lack of a maltotriose transporter on the cell surface since maltose and maltotriose are both hydrolyzed by the same α-glucosidase.
Potential Uses in Brewing
Implementation of S. eubayanus as a brewing yeast is not practical in the immediate future for commercial breweries. There are too many unknowns with how the yeast performs and to how to best utilize it. There is, however, an opportunity to make new S. pastorianus strains. As GM yeast strains are not an option in the commercial production of beer, breeding S. cerevisiae strains that have a desirable phenotype with S. eubayanus with another desirable phenotype may produce lager strains that perform better than the current strains in use.
Homebrewers, on the other hand, have a unique opportunity to characterize a new yeast species in brewing. While a few academic studies have analyzed fermentation of wort by S. eubayanus, there is very little information on the quality of a beer produced by this yeast. Since S. eubayanus is not yet domesticated for brewing, reusing the yeast over successive generations may select for a better fermenting yeast. In addition, the more the fermentation profile of this yeast is characterized, the better recipes can be formulated around it.
There is a lot to learn about S. eubayanus, and the next several years may prove to be a very exciting time learning how to best utilize this yeast in the brewhouse.
References:
1. Corran, H. S. A History of Brewing. 1975. London, UK, David and Charles.
2. Libkind D, Hittinger CT, Valério E, Gonçalves C, Dover J, Johnston M, Gonçalves P, Sampaio JP (2011) Microbe domestication and the identification of the wild genetic stock of lager-brewing yeast. Proc Natl Acad Sci U S A. Aug 30;108(35):14539-44
3. Peris D, Sylvester K, Libkind D, Gonçalves P, Sampaio JP, Alexander WG, Hittinger CT (2014) Population structure and reticulate evolution of Saccharomyces eubayanus and its lager-brewing hybrids. Mol Ecol. Apr;23(8):2031-45.
4. Bing J, Han PJ, Liu WQ, Wang QM, Bai FY (2014) Evidence for a Far East Asian origin of lager beer yeast. Curr Biol. May 19;24(10):R380-1.
5. González SS, Barrio E, Gafner J, Querol A (2006) Natural hybrids from Saccharomyces cerevisiae, Saccharomyces bayanus and Saccharomyces kudriavzevii in wine fermentations. FEMS Yeast Res. Dec;6(8):1221-34.
6. Lindner P. (1909) Mikroskopische Betriebskontrolle in den Garungswerben (6th edn). Berlin: Paul Parey.
7. Guilliermond A (1920) The Yeasts. (FW Tanner, Trans). New York: John Wiley and Sons, Inc.
8. Dunn B, Sherlock G (2008) Reconstruction of the genome origins and evolution of the hybrid lager yeast Saccharomyces pastorianus. Genome Res. Oct;18(10):1610-23.
9. Nakao Y, Kanamori T, Itoh T, Kodama Y, Rainieri S, Nakamura N, Shimonaga T, Hattori M, Ashikari T (2009) Genome sequence of the lager brewing yeast, an interspecies hybrid. DNA Res. Apr;16(2):115-29.
10. Walther A, Hesselbart A, Wendland J (2014) Genome sequence of Saccharomyces carlsbergensis, the world’s first pure culture lager yeast. G3 (Bethesda). Feb 27;4(5):783-93.
11. Hewitt SK, Donaldson IJ, Lovell SC, Delneri D (2014) Sequencing and characterisation of rearrangements in three S. pastorianus strains reveals the presence of chimeric genes and gives evidence of breakpoint reuse. PLoS One. Mar 18;9(3):e92203.
12. Gibson BR, Storgårds E, Krogerus K, Vidgren V (2013) Comparative physiology and fermentation performance of Saaz and Frohberg lager yeast strains and the parental species Saccharomyces eubayanus. Yeast. Jul;30(7):255-66.